NA is synthesized in each NB and PCC tissues, and various members of this synthetic pathway have been reported previously in NB as best scoring classifiers. The significance of this pathway is very well characterized in NB, nevertheless, the expression of numerous of its important mem bers have not been reported in PCC, yet. The homeobox transcription aspect PHOX2B is essen tial for your differentiation and servicing of noradre nergic neurons. PHOX2B mutations are observed in uncommon hereditary kinds of NB and Hirschsprungs disorder. PHOX2B is concerned during the transcriptional regula tion of RET expression, nevertheless mutations of PHOX2B in hereditary and sporadic PCC have not been reported, nevertheless. The downstream targets with the master regulators of neural crest derived precursor cell growth PHOX2B and HAND2 are PHOX2A, GATA2, GATA3.
These transcription elements perform critical roles from the expressional selleckchem regulation in the TH and DBH genes. TH catalyzes the charge limiting phase of NA production, and DBH converts dopamine to NA. NA in chromaffin cells is co stored in secretory granules with CGA and CGB molecules. The uptake into granules is mediated by SLC18A1 protein through the cytoplasm and by SLC6A2 from your extracellular area. From the comparison of NB or PCC tissues with other tumors and typical tissues, we’ve got discovered the signifi cant overexpression of all the above stated genes in NB or PCC samples in greater than 90 comparisons. This observation highlights the common origin of NB and PCC, nonetheless, it really is surprising that you’ll find only extremely number of data over the expression of those genes in PCC.
Similarities amongst NB and PCC To investigate probably the most major similarities amongst NB and PCC tissues, we’ve searched for previously unreported reference genes. By this strategy, by far the most equivalent pathway among NB and PCC was death buy Ibrutinib recep tor signaling. Avoidance of apoptosis is really a vital method of tumorigen esis. Considering the fact that this pathway included essentially the most similar gene expression patterns in NB and PCC, we could conclude that these tumors may use the very same techniques for rescue from programmed cell death. The importance of proteins involved from the regulation of apoptosis triggered by death receptors is already reported in NB. Apoptosis in mammalian cells could be initiated as a result of two major pathways, one particular in volving tumor necrosis aspect alpha and DR1 6, another involving release of cytochrome c from mito chondria as a result of Fas activation.
Numerous reports underline the significance of mitochondria dependent signaling in NB. Fas resistance in NB could create through the inactivation of caspase 8, which can be absent in extra 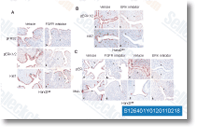 than 1 third of NB cases, and normally methylated in over 60% of PCC. Fas resistance may well even build in caspase eight expressing NB by means of large level expression on the antiapoptotic protein BCL2.
than 1 third of NB cases, and normally methylated in over 60% of PCC. Fas resistance may well even build in caspase eight expressing NB by means of large level expression on the antiapoptotic protein BCL2.
Monthly Archives: May 2014
Here, we presented our efforts to create a modeling framework f
Right here, we presented our efforts to develop a modeling framework for constructing large scale kinetic models that mechanis tically hyperlink transcriptional regulation and metabolism. This permitted us to gain knowing of complicated physiological relations from fluxome, metabolome, and gene expression data. We demonstrated the capacity of our technique to cap ture these relations, its versatility to simulate unique ex periments, and its robustness with respect to modeling approximations and information uncertainty by analyzing the re sponse of S. cerevisiae beneath unique pressure situations. Importantly, our strategy can be utilized to other orga nisms of healthcare and industrial relevance for which a metabolic network reconstruction, metabolic flux measurements, and gene expression information can be found for that problems of interest.
The system offers productive remedies to massive scale modeling challenges On the list of big issues in constructing large scale kinetic versions could be the definition of proper reaction fee expressions. Rather inhibitor NVP-AUY922 of defining mechanistic response fee expressions on a situation by case basis, some approaches streamline this approach by relying on generic expressions to translate a metabolic network right into a kinetic model in an automated or semi automated vogue. Unique gen eral forms are actually proposed, such as log linear kinetics, Michaelis Menten form kinetics, convenience kinetics, or GMA kinetics. GMA kinetics are employed, as an example, in ensemble modeling and mass action stoichiometric simulation designs.
AG014699 In ensemble modeling and MASS designs, the enzymatic reactions are decomposed into their elementary ways, and every stage is then modeled employing mass action kinetics. The decomposition increases the resolution with the model, preserves enzyme saturation conduct, and simplifies the parameter estimation issue, but in the rate of consid erably growing the dimension in the model and also the volume of data required to estimate parameter values. In contrast, we used a special situation of GMA kinetics that involves a minimum quantity of parameters, which might be obtained immediately from out there experimental data. Moreover, enzymatic reactions weren’t decomposed into elementary measures in order to avoid in creasing the dimension from the model. A further challenge is definitely the determination of model par ameter values. The problems in solving this trouble is linked to the type of the kinetic expressions and to the availability of experimental information.
If experimental data are usually not offered, approaches such as log linear kinetics and comfort kinetics demand mining the literature for parameter  values, which could be impractical for substantial scale models. Approaches working with GMA kinetics partially stay clear of literature mining. In these approaches, such as MASS modeling, thermodynamic information and facts collected from the litera ture is mixed with experimentally determined me tabolite and/or enzyme concentrations and flux distribu tions to estimate the remaining model parameters.
values, which could be impractical for substantial scale models. Approaches working with GMA kinetics partially stay clear of literature mining. In these approaches, such as MASS modeling, thermodynamic information and facts collected from the litera ture is mixed with experimentally determined me tabolite and/or enzyme concentrations and flux distribu tions to estimate the remaining model parameters.
Usually aromatic hydrocarbons are a lot more recalci trant than a
On the whole aromatic hydrocarbons are more recalci trant than aliphatic hydrocarbons to microbial degrad ation. The Troll samples all share the common predominant source of hydrocarbons, the underlying oil and gas reservoir. The improved genetic potential for degradation of aromatic hydrocarbons in Tplain and Tpm1 two is as a result likely to be a result of sequential degradation in the numerous fractions in oil. A more energetic subcommunity in Tplain and Tpm1 two could have degraded a larger fraction from the significantly less recalcitrant aliphates, forcing a shift from the metabolism in the direction of greater degradation of aromatic hydrocarbons at the sampling time. The seabed is often a dynamic environment, in addition to a concept by Hovland and coworkers proposes that as previous pockmarks are closed down new ones are produced as a result of adjustments in fluid flow pathways in excess of time.
Higher likely for hydrocarbon degradation, potentially inhibitor ONX-0914 relevant to a extra active hydrocarbonoclastic subcommunity in Tplain and Tpm1 2, may very well be explained by enhanced bioavailability of critical nutrients and metals involved in hydrocarbon deg radation at these sites in contrast to your other Troll websites, like a consequence of greater porewater seepage. Enhanced porewater seepage could also bring about a slightly greater hydrocarbon availability, specifically on the much more aqueous soluble hydrocarbons, which could sustain a more active hydrocarbonoclastic subcommunity at Tplain and Tpm1 two. At Tpm1 2 a potential boost in porewater seepage could possibly be explained from the carbonate mound identified near to the sampling site. This carbonate mound could constitute a seal for fuel migrating in the direction of the seafloor, thereby escalating the strain within the porewater forced out along its sides. Additional, variations in publicity to water present activ ity could also have an effect on the bioavilibility of nutrients and community structure.
Preceding investigation selleck of fauna in significant Troll pockmarks has indicated the probability for improved currents or turbulence at the eastern slope in the pockmarks within the region. Likewise, there exists no protection in the water present to the Troll plain. Methane oxidation in pockmark sediments Despite the fact that methanotrophs contributed to all seven meta genomes, no basic overabundance can be detected during the Troll pockmark metagenomes in contrast to your Oslofjord metagenomes, supporting the geochemical conclusion that there is no, or quite low, lively methane seepage in these pockmarks with the current time. We did understand marker genes for aerobic methane oxidation in Tpm1 two and Tplain. This might be connected to your slight overabundance of aerobic methanotrophic taxa in these samples. Interestingly, reads associated with ANME were two to 3 times much less abundant from the metagenome through the Troll plain, than from the Troll pockmark metagenomes wherever ANME accounted for up to 0.
In these research sequence homologues of all enzymes wanted for C
In these research sequence homologues of all enzymes needed for CO2 based methanogenesis with exception of N5, N10 methylene tetrahydromethanop terin reductase had been recognized. Methyl coenzyme M reductase is assumed to catalyze the first stage of AOM as well as final phase of methanogenesis, and is hence a marker gene for each processes. Similarly, dissimilatory sulphite reductase is usually employed being a marker gene for SRB. When oxygen is current, aerobic methanotrophs are lively in methane oxidation. Identified aerobic methano trophs include things like representatives of Gammaproteobacteria, Alphaproteobacteria and Verrucomicrobia. These organisms convert methane to methanol applying the enzyme methane monooxygenase. The particu late, membrane bound model of methane monooxygen ase, found in all aerobic methanotrophs, is employed like a marker gene for aerobic oxidation of methane.
The methanol formed is converted to formaldehyde, and that is assimi lated by considered one of two known pathways. Sort I and sort II methanotrophs employ the ribulose monophosphate pathway as well as serine pathway respectively. Variety ? methanotrophs use principally the ribulose monopho sphate pathway, but possess the enzymes needed for your serine pathway as well. Secure isotope probing and sequencing of 16S kinase inhibitor Dinaciclib rDNA and pmoA, too as lipid biomarker evaluation, have detected variety I aerobic methanotrophs in sediments and biofilms on the COP Shane and Brian seeps. Not too long ago, measurements of common 13C of carbonates and lipid biomarkers connected with ANME and SRB also indicated occurrence of AOM with the Brian seep. Yet another survey in the Brian seep detected ANME two at six 9 cm bsf by FISH. While in the current review, we’ve got used metagenomics to characterize the taxonomic and metabolic potential for the two aerobic and anaerobic methane oxidation in two sediment samples from distinct depths with the Tonya seep.
By steering clear of PCR amplification and primer target specificity, the metagenomics strategy LY2109761 presented even further insight in to the taxonomy and metabolic poten tial on the prokaryotic communities in the methane seep sediments. Final results Fuel measurements and methane oxidation rate The average methane oxidation fee based on eleven mea surements within the prime 15 cm of your seep sediments was 156 64 nmol cm three day one. Nonetheless, the fuel emitted through the Tonya seep sediments into the water phase con tained a significant fraction of methane. Even after travelling 25 m through the water column, where dissolved O2 and N2 entered the bubbles, the two fuel samples con tained 80. 4% and 68. 1% methane. When O2 and N2 have been excluded, and the hydrocarbon and CO2 content had been normalized, methane accounted for 93. 6% in the two gasoline samples. The remainder consisted of CO2 and short chain hydrocar bons.
The sample for RNA sequencing was derived from pooling of the R
The sample for RNA sequencing was derived from pooling in the RNA samples isolated in the unique tissues in accordance to your following ratios. two roots.1 pseu dostems.1 leaves.1 fruits.1 flowers. RNA processing for transcriptome sequencing Poly enriched mRNA was purified from your complete RNA samples using Sera mega Oligo beads and fragmented with divalent cations at elevated temperature. The RNA fragments have been applied for cDNA synthesis by using the SuperScript cDNA synthesis kit with random hexamer primers, Following finish repairing, cDNA fragments have been ligated to adaptors, purified and PCR amplified to generate the library which was then sequenced making use of Illumina HiSeq 2000. RNA processing for digital gene expression examination The tag libraries were prepared applying the NlaIII sample prep kit according to your.
Following mRNA enrichment buy inhibitor and cDNA synthesis as described over, five ends of tags were gener ated by digesting with NlaIII, The fragments aside from the three cDNA fragments linked to Oligo beads were washed away and the Illumina adaptor 1 was li gated to your sticky 5 end of the digested bead bound cDNA fragments. in the DNA fragments have been minimize with MmeI. Right after removing 3 fragments with magnetic beads precipitation, Illumina adaptor two was ligated on the 3 ends of tags. The adaptor ligated cDNA tags were enriched by 15 cycles of linear PCR amplification as well as resulting 85 bp fragments were purified from 6% acrylamide gel. Right after denaturing, the single chain mole cules had been fixed onto the Illumina Sequencing Chip for sequencing.
Transcriptome assembly and evaluation from RNA seq The raw reads have been cleaned the full report by removing adaptor se quences and low quality reads with ambiguous N. TopHat, a splice junction mapper for RNA Seq reads, was utilized to align RNA seq reads on the Musa genome sequence with default parameters, Cufflinks was then applied to assemble the transcripts through the TopHat alignment benefits. Novel genes have been recognized by comparing all the assembled transcripts to banana genome annotation by Cuffcompare within the cufflinks package. The novel loci located by Cufflinks have been scanned for ORF by coding annotation instrument in Trinity package deal, People transcripts with a putative full ORF have been aligned to the NCBI nr database as well as the Uni Prot plant protein sequences by BLASTx to locate homologous proteins.
The transcripts with extra than one exon or single  exon but acquiring hits to acknowledged proteins at E value cutoff 1e 5 had been reported as final novel transcripts despite the fact that some of the other sequences could also derived from genes which have not been annotated. Identification of SNPs and indels SAMtools was made use of to analyze the achievable SNPs and indels during the banana genome according to the transcrip tome data. The authentic reads were mapped back for the assembled banana transcripts. The SNPs and indels were referred to as employing the mpileup instrument in SAMtools bundle.
exon but acquiring hits to acknowledged proteins at E value cutoff 1e 5 had been reported as final novel transcripts despite the fact that some of the other sequences could also derived from genes which have not been annotated. Identification of SNPs and indels SAMtools was made use of to analyze the achievable SNPs and indels during the banana genome according to the transcrip tome data. The authentic reads were mapped back for the assembled banana transcripts. The SNPs and indels were referred to as employing the mpileup instrument in SAMtools bundle.
The sample for RNA sequencing was derived from pooling of your
The sample for RNA sequencing was derived from pooling of your RNA samples isolated from the different tissues according on the following ratios. two roots.1 pseu dostems.1 leaves.one fruits.one flowers. RNA processing for transcriptome sequencing Poly enriched mRNA was purified in the total RNA samples utilizing Sera mega Oligo beads and fragmented with divalent cations at elevated temperature. The RNA fragments have been applied for cDNA synthesis through the use of the SuperScript cDNA synthesis kit with random hexamer primers, Immediately after finish repairing, cDNA fragments had been ligated to adaptors, purified and PCR amplified to generate the library which was then sequenced applying Illumina HiSeq 2000. RNA processing for digital gene expression analysis The tag libraries have been ready applying the NlaIII sample prep kit in accordance to your.
Following mRNA enrichment this article and cDNA synthesis as described over, 5 ends of tags have been gener ated by digesting with NlaIII, The fragments apart from the 3 cDNA fragments linked to Oligo beads have been washed away and the Illumina adaptor one was li gated on the sticky 5 end of your digested bead bound cDNA fragments. of your DNA fragments had been lower with MmeI. After removing three fragments with magnetic beads precipitation, Illumina adaptor 2 was ligated for the 3 ends of tags. The adaptor ligated cDNA tags had been enriched by 15 cycles of linear PCR amplification as well as resulting 85 bp fragments were purified from 6% acrylamide gel. Just after denaturing, the single chain mole cules have been fixed onto the Illumina Sequencing Chip for sequencing.
Transcriptome assembly and examination from RNA seq The raw reads were cleaned additional resources by removing adaptor se quences and minimal quality reads with ambiguous N. TopHat, a splice junction mapper for RNA Seq reads, was employed to align RNA seq reads for the Musa genome sequence with default parameters, Cufflinks was then employed to assemble the transcripts from your TopHat alignment success. Novel genes have been identified by comparing each of the assembled transcripts to banana genome annotation by Cuffcompare in the cufflinks package deal. The novel loci discovered by Cufflinks had been scanned for ORF by coding annotation tool in Trinity package, Individuals transcripts that has a putative full ORF were aligned to the NCBI nr database and the Uni Prot plant protein sequences by BLASTx to find homologous proteins.
The transcripts with much more than one particular exon or single  exon but acquiring hits to regarded proteins at E value cutoff 1e 5 have been reported as final novel transcripts although some of the other sequences could also derived from genes which have not been annotated. Identification of SNPs and indels SAMtools was employed to analyze the possible SNPs and indels while in the banana genome according to the transcrip tome data. The original reads had been mapped back towards the assembled banana transcripts. The SNPs and indels were known as employing the mpileup tool in SAMtools package.
exon but acquiring hits to regarded proteins at E value cutoff 1e 5 have been reported as final novel transcripts although some of the other sequences could also derived from genes which have not been annotated. Identification of SNPs and indels SAMtools was employed to analyze the possible SNPs and indels while in the banana genome according to the transcrip tome data. The original reads had been mapped back towards the assembled banana transcripts. The SNPs and indels were known as employing the mpileup tool in SAMtools package.
SP07 is often a marked exception inside the latter regard, A fu
SP07 can be a marked exception during the latter regard, A different oddity amid these sequences is four of them are truncated C terminally with halt codons, in spite of the truth that SP01 and 07 show expression levels of 9. 6 and 7. 1%, respectively. Wang et al. reported that a Kentucky population of Crotalus horridus lacks an acidic PLA2 for the reason that the codon for Tyr22 has mutated into a prevent codon. They concluded that minimal PLA2 expression amounts in most Crotalus horridus venoms is often attributed to translation blockage. At this time, it truly is complicated to learn how wide spread this phenomenon might be, but it is apparent that these two Ovophis SPs are translated successfully given that they’d ample peptide coverage, L amino acid oxidase The Protobothrops transcriptome incorporated two transcripts for L amino acid oxidase, compris ing 2.
3% and 6. 8% of all transcripts, respectively, A single LAO transcript was existing in Ovophis glands, representing 0. 6% on the transcriptome, Peptides accounting for 84. 6% and 70. 8% of Protobothrops LAO 1 and LAO 2, respectively, and 78. kinase inhibitor pf-562271 7% of the Ovophis LAO transcript sequence was identified by mass spectrometry, Small venom constituents Cysteine wealthy secretory proteins Two CRISPs had been identified in the Protobothrops transcrip tome, CRISP 1, for which a finish transcript was obtained, is identical to triflin, but CRISP 2 aligns finest by using a CRISP bearing an EGF like calcium binding domain through the venom of Crotalus adamanteus, Nonetheless, the putative 39 residue EGF domain during the C. adamanteus toxin isn’t going to align very well with the corresponding region of your Protobothrops transcript.
The latter includes only 4 acidic residues, Odanacatib in contrast with nine in the C. adamanteus sequence. Only 3 from the five C. adamanteus cysteine residues match, as well as two sequences require a two residue gap to attain even this poor alignment. Therefore, we assume it unlikely that there’s a functional EGF like calcium binding domain while in the Protobothrops toxin. Additionally, no peptides were sequenced for this odd CRISP, whereas 84. 6% of CRISP one was sequenced. A single, comprehensive CRISP transcript was recognized within the Ovophis transcriptome, but sequenced peptides accounted for 89. 0% of its primary structure. It had been most similar to a CRISP from the venom of Bothriechis schlegelii, CRISPs are normally not abundant components of snake venoms, however they are extensively distributed taxonomically.
Ablomin, triflin and latisemin are L variety Ca2 channel antagonists of depolarization induced arterial smooth muscle 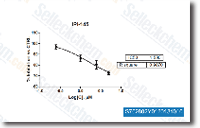 contraction, nevertheless they never affect caffeine induced contraction, as a result they advertise vasodilation and hypotension. Tigrin from venom with the Japanese colubrid, Rhabdophis tigrinus, impacted neither. This is often almost certainly mainly because Rhabdophis venom glands are not secretory in nature.
contraction, nevertheless they never affect caffeine induced contraction, as a result they advertise vasodilation and hypotension. Tigrin from venom with the Japanese colubrid, Rhabdophis tigrinus, impacted neither. This is often almost certainly mainly because Rhabdophis venom glands are not secretory in nature.
Flusilazole remedy decreased the expression of genes concerned wi
Flusilazole treatment method decreased the expression of genes concerned with PSI, PSII action and photosynthetic electron transport in photosystem. The down regulated genes included individuals that encode subunit V, PsaA PsaB, F variety H transporting ATPase subunit gamma in photosystem I, and 13 kDa protein in photosystem II, We identified that GA3 remedy led to sturdy induction in photosynthesis connected gene expression, These final results indicate that photosynthesis may play a vital purpose in the calyx abscission and samples with diminished photosynthesis perform are more vulnerable to abscission. Plant hormone signal transduction Many sorts of hormones regulate the course of action of abscis sion, such as IAA, ABA, GA, JA, and ethylene, amongst which IAA and ethylene play a pivotal position.
IAA pre vents, when ethylene accelerates the abscission processes, Forty three genes have been noticed to be concerned in ethylene synthesis, perception, and response within the current examine. ACC synthase and ACC oxidase are fee limiting enzymes in ethylene biosyn thesis, Genes concerned in ACC synthase showed expressions in creased by 2. 109 to six. 236 times through the early calyx abscission selleck inhibitor process immediately after Flusilazole treatment method. Two ACC oxidase genes were up regulated in later calyx abscission procedure immediately after Flusilazole remedy.
Then again, two ethylene receptor genes and 3 ethylene responsive genes have been up regulated in later selleck chemical Lonafarnib calyx ab scission method right after Flusilazole treatment, That is consistent with all the correlation involving abscission and enhanced expression of genes for ethylene synthesis and ethylene receptors in AZ, which is reported in mature fruit of olive and apple, The general rule states that offered the flux of IAA towards the abscission zone is maintained, cell separation is inhibited and abscission won’t transpire, A single hundred and two auxin related genes have been differentially expressed from the Flusilazole and GA3 deal with ments, including indole three acetic acid amido synthetase, indole three acetic acid induced protein, and auxin response element. Nine genes encoding AUX IAA protein have been down regulated at 6 d right after Flusilazole remedy and 13 genes encoding auxin responsive aspects have been also down regulated at 10 d soon after Flusilazole treatment. Genes relevant to polar auxin transporting have been also impacted by GA3 treatment method, with auxin influx carriers induced and efflux carriers largely repressed.
We noticed that an auxin transport associated gene was up regulated at six d, but repressed at 10 d right after Flusilazole therapy. Moreover, a gene encoding spermidine synthase was up regulated with Flusilazole remedy at onset of calyx abscission professional cesses. As talked about above, hormones seem to perform a reasonably important part through the early phases of abscission within the calyx, considering that a vast majority from the transcrip tionally activated aspects involved in hormone signaling seem to be downstream of the induction of abscission.
Every single chip contained four repetitions of every probe In t
Each chip contained four repetitions of each probe. In complete, the 1,215 miRNAs have been composed of 224, 496, 148 and 347 miRNAs from Arabidopsis thaliana, Oryza sativa, Sorghum bicolor and Zea mays, individually. RNA labeling, microarray hybridization, array scanning, and datas examination were performed basically as previously described, Bioinformatics examination of sequencing information The two little RNA reads and degradome reads were gener ated by Illumina Genome Analyzer II. As to the minor RNA library, the data have been processed and analyzed as pre viously described by Wang et al. and Zhang et al, In brief, different reads ranging from18 25 nt have been col lected and mapped towards the maize genome reference sequences by SOAP2, Soon after getting rid of sequences matching non coding rRNAs, tRNAs, snRNAs and snoRNAs while in the Rfam and NCBI Genbank databases, the matched Solexa reads that had been extracted 250 nt in the sequence flanking the genomic sequences were employed for RNA secondary framework prediction, which was performed by mFold 3.
5 and analyzed by MIREAP to determine new candidates employing default settings. The candidate miRNA checklist was even further trimmed based on the criteria as described, Based mostly within the hairpin construction of your pre miRNA, the corresponding miRNA star sequence was also identified. Degradome reads had been filtered applying customized Perl script. The remaining distinct 20 21 nt sequences that properly matched selleck chemicals Epigenetic inhibitor maize contigs had been collected for further evaluation.
The 15 nt upstream and 5 end on the reads that mapped to maize contigs were extracted to create 30 sequence tags, which had been made use of to align to newly identified miRNAs and miRBase employing the Cleave and pipeline, Alignments had been collected as candidate Carfilzomib targets when they fulfilled the criteria as described ahead of, GO functional enrichment evaluation of all candidate tar gets while in distinctive developmental stages was carried out making use of Blast2GO and GO annotations have been performed applying AgriGO, KEGG pathway analyses of differentially expressed genes were performed making use of Cytos cape program together with the ClueGO plugin, Stem loop quantitative true time PCR analysis Validations of 13 randomly selected mature miRNAs were carried out by stem loop reverse transcription PCR, Total RNA was applied to initiate the reverse transcription response. Primers for that stem loop RT PCR have been constructed making use of approaches as de scribed by Chen et al. and Varkonyi Gasic et al, The stem loop RT PCR was working with the Applied Bio methods 7500 Actual Time PCR Procedure, All primers have been listed in Supplemental file twelve. Table S8. All reactions were run in triplicate. 5S rRNAs was utilised because the internal handle for stem loop RT PCR, The RNA sequencing information have been deposited from the NCBI beneath the accession number GSE47837.
Complete RNA samples with 28S 18S ratios in a vary from one 8 to
Total RNA samples with 28S 18S ratios within a range from 1. 8 to two. 0 and RNA integrity index from eight. 0 to 10. 0 have been picked for even more processing. PolyA RNA fraction was purified from total RNAs using a MACS mRNA isolation kit, Double stranded cDNA was synthesized making use of a Superscript cDNA syn thesis kit with random hexamer primers at a concentration of 5 uM. The cDNA fragments had been sheared making use of a Covaris E110 for 75 sec below problems Duty cycle of 20% and Intensity of five, then the cDNA fraction with lengths of 200 250 bp was excised from 8% polyacryl amide gel electrophoresis for cDNA library con struction applying a paired end sample prep kit, Briefly, the cDNAs had been topic to end restore and phosphorylation by T4 DNA polymer ase, Klenow DNA polymerase, and T4 polynucleotide kinase respectively in a single response.
The cDNAs with 3 A overhangs, generated by Klenow fragment, had been ligated to paired finish adapters, which have 5 T overhangs. The adapter ligated products were purified and enriched by PCR for 10 to 15 cycles utilizing Phusion DNA polymerase and paired finish primer selleck tsa inhibitor set, PCR product or service of your preferred dimension selection was purified using 8% Page. The purified cDNA quality was assessed and quantified applying an Agilent DNA 1000 series II assay kit on the Agilent 2100 Bioanalyzer plus a Quant IT ds DNA HS assay kit around the Qubit fluorometer, The cDNA library was diluted to 8 nM and this final concentration was checked and determined again by Quant IT dsDNA HS Assay before Illumina sequencing.
Illumina RNA seq analysis The Illumina GA IIx platform was utilised for deep sequen cing from the cDNA libraries from both five and three end for 76 bp reads following the makers guide at the British Columbia Cancer Company, The deconvolution of supplier Dinaciclib fluorescent images to DNA sequences, base calling and high-quality worth calcula tion had been performed using the Illumina data processing pipeline, The raw Illumina 76 bp pair end sequences were deposited from the NCBI Sequence Read Archive beneath accession numbers SRR1013833, SRR1013836, and SRR1013837. Bioinformatics analyses The CLC genomics workbench was picked for de novo transcrip tome assembly from the existing review because the CLC software package includes a a lot quicker computing pace with comparable or improved assembly benefits than other bioinformatics professional grams, Reads and read stretches of bad high-quality bases have been eliminated and trimmed with filter threshold at high quality score, ambiguous nucleotide, and length trimming just before de novo transcrip tome assembly.
Making use of the BLASTn and tBLASTx algorithms, all non redundant contigs had been utilised for a BLAST search against the NCBI nr database, the PGI or SGI database, a Melampsora laricis popu lina protein database, along with a set of P. monticola ESTs, The PGI and SGI databases contained 77,326 and 79,409 exceptional ESTs respectively.
